Cytosplore
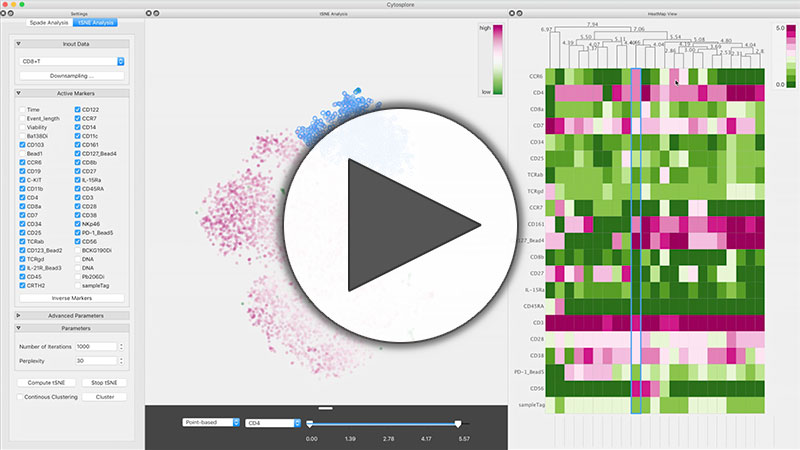
Cytosplore is an interactive visual analysis system for understanding how the immune system works. The goal of the analysis framework is to provide a clear picture of the immune systems cellular composition and the cells’ corresponding properties and functionality. Cytosplore is targeted at the analysis of mass cytometry (CyTOF) data. Mass cytometry is a novel technique to determine the properties of single-cells with unprecedented detail. This amount of detail allows for much finer differentiation but also comes at the cost of more complex analysis. Cytosplore incorporates state-of-the art clustering and dimensionality reduction techniques. The inclusion of HSNE allows the exploration of millions of cells without the need for downsampling. Furthermore Cytosplore provides Approximated-tSNE (up to 100x faster than standard t-SNE without loss in precision) and a custom implementation of the SPADE clustering algorithm. Cytosplore implements progressive visual analytics and visualization techniques to provide a highly engaging user experience through direct feedback and linked views. Try Cytosplore yourself and sign up for our mailing list for latest news.
Team
TU Delft / LUMC
Lead Developer/Project Liaison
Philips, former TU Delft
Algorithms Research
Stanford University
Immunology Research
LUMC
Development
LUMC, former TU Delft
Development
LUMC
PI Immunology
TU Delft
PI CGV
LUMC
PI LKEB
TU Eindhoven
PI VAnPIRe
Join the team! We have open projects within the Cytosplore project including a project on classification of cellular expression data through dimensionality reduction. If you are interested in working with us on this or other projects involving biological visualization for a research project at the TU Delft, LUMC, or TU Eindhoven, please contact Thomas Höllt or Anna Vilanova to discuss the possibilities.
Publications & Media
Media
The Delta published an article covering our latest results including HSNE in November 2017: New tool identifies a few cells among millions
Press releases by TU Delft and LUMC covering the publication of our Nature Communications paper.
The Cytosplore team was interviewed for an article in Medical Delta in December 2016: Exploring new dimensions of the immune system.
Selected Publications
Cytosplore: Interactive Immune Cell Phenotyping for Large Single-Cell Datasets. Computer Graphics Forum, 2016.
Abstract: In this work, we present Cytosplore, implementing an interactive workflow to analyze mass cytometry data in an integrated system, providing multiple linked views, showing different levels of detail and enabling the rapid definition of known and unknown cell types, based on mass cytometry (CyTOF). Cytosplore handles millions of cells, each represented as a high-dimensional data point, facilitates hypothesis generation and confirmation, and provides a significant speed up of the current workflow.
Visual Analysis of Mass Cytometry Data by Hierarchical Stochastic Neighbor Embedding Reveals Rare Cell Types. Nature Communications, 2017.
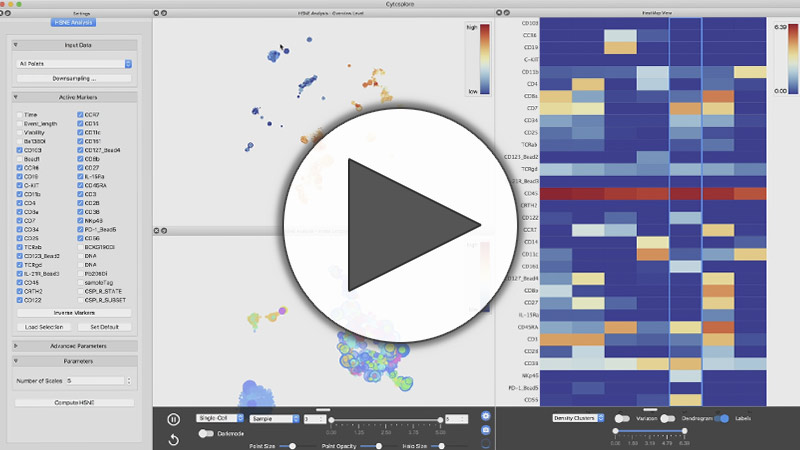
Abstract: In this work, we present Cytosplore+HSNE. HSNE constructs a hierarchy of non-linear similarities that can be interactively explored with a stepwise increase in detail up to the single-cell level. We applied HSNE to several available mass cytometry datasets. We found that HSNE efficiently replicates previous observations and identifies rare cell populations that were previously missed due to downsampling in a fraction of the time.
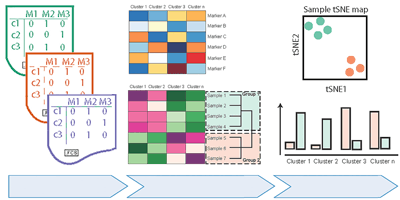
Cytofast: A workflow for visual and quantitative analysis of flow and mass cytometry data to discover immune signatures and correlations. Computational and Structural Biotechnology Journal, 2018.
Abstract: Cytosplore incorporates clustering and dimensionality reduction techniques and algorithms allowing visualization of multi-parametric cytometric analysis. To additionally provide means to quantify specific cell clusters and correlations between samples, we developed an R-package, called cytofast, for further downstream analysis. Cytofast enables the visualization and quantification of cell clusters for an efficient discovery of cell populations associated with diseases or physiology.
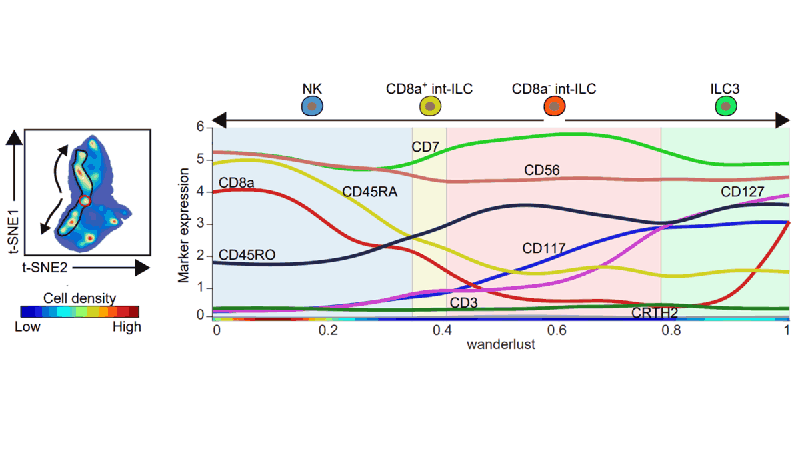
Mass Cytometry Reveals Innate Lymphoid Cell Differentiation Pathways in the Human Fetal Intestine. Journal of Experimental Medicine, 2018.
Abstract: Innate lymphoid cells (ILCs) are abundant in mucosal tissues and involved in tissue homeostasis and barrier function. We applied mass cytometry to analyze ILCs in the human fetal intestine and distinguished 34 distinct clusters through a t-SNE-based analysis. By visualizing the dynamics of the t-SNE computation, we identified smooth phenotypic transitions from cells within the Lin-CD7+CD127-CD45RO+CD56+ cluster to both the NK cells and CD127+ ILCs...
Get Cytosplore
Disclaimer: Cytosplore is a research project between TU Delft and Leiden University Medical Center. It is not a commercial software product and not licensed for clinical use. We do our best to provide support in our free time but cannot guarantee it. If you have any suggestions, problems, want to share your success stories or papers published using Cytosplore, we would love to get feedback! Please do not hesitate to get in touch.
Cytosplore is free for academic and non-commercial use, however, if you use Cytosplore within the scope of a scientific article you must cite the original publications:
T. Höllt, N. Pezzotti, V. van Unen, F. Koning, E. Eisemann, B. Lelieveldt, and A. Vilanova. Cytosplore: Interactive Immune Cell Phenotyping for Large Single-Cell Datasets. Computer Graphics Forum (Proceedings of EuroVis), 35(3): pp. 171—180, 2016.
V. van Unen, T, Höllt, N. Pezzotti, N. Li, M. Reinders, E. Eisemann, A. Vilanova, F. Koning, and B. Lelieveldt. Visual Analysis of Mass Cytometry Data by Hierarchical Stochastic Neighbor Embedding Reveals Rare Cell Types. Nature Communications, 2017
For other operating systems or older versions please see the release archive.
Cytosplore Mailinglist
We will use the information you provide on this form to be in touch with you and to provide updates on Cytosplore Development. You can change your mind at any time by clicking the unsubscribe link in the footer of any email you receive from us, or by contacting us at [email protected]. By clicking below, you agree that we may process your information in accordance with these terms.
We use MailChimp as our mailing automation platform. By clicking below to submit this form, you acknowledge that the information you provide will be transferred to MailChimp for processing in accordance with their Privacy Policy and Terms.